With my previous blog post about scoping rules (check it out here!) turning out to be a popular success I wanted to take scoping rules one step further – to applications! And what is the easiest and fastest way to make an interactive application for a user in R? You guessed it! SHINY!
For those not familiar with using Shiny to create applications there is good documentation on RStudio’s website with the start of a showcase gallery as well. I’d suggest you check out the documentation and give your first Shiny application a try – you’ll be hooked! If a picture is worth 1000 words, an interactive application is priceless! Especially when you can give your customer, users, boss, etc a great experience around the analytics work you have spent so much time on! The ooh-ahh factor is not just nice-to-have, it’s really essential to making sure your audience understands your results!
Enough about why you should try out Shiny! For initial users of Shiny creating an application is a generally straightforward process to get good results. The great part about this is that you don’t need to worry about scoping right away since you are the only user of your application. However, once you start creating more applications in Shiny you will start to deploy them to a Shiny server, and have (potentially) more than one user per application and you certainly don’t want your users colliding with each other! This is when scoping becomes critical!
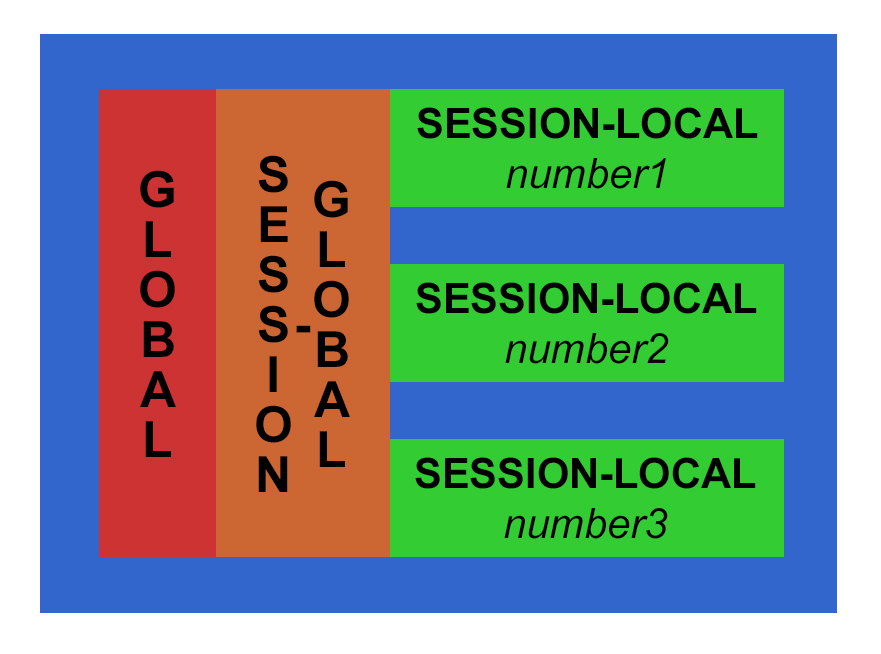
There are three main scopes in a standard shiny application – the global, session-global and session-local scopes.
The global scope, which is the easiest to understand (and most dangerous to use)! Anything declared within the global scope is accessible to both the UI and Server sections of the application as well as inside and outside the session scope. The only way to declare a globally-scoped variable in a shiny application that is not a single file is to use the global.R file. Luckily this makes it obvious what scope anything in this file is in!
The next most common scope is session-global. This is a nearly-global scope and in a standard two-file (session.R and ui.R) shiny application this is the most (or closest to true) global scope you will encounter. In this scope declared items are visible to all R server sessions. There is no differentiation between users; all users receive, can see, and can use or change these items. The most appropriate reason to use this scope is for items that are globally relevant and generally do not change based on users or their input, such as reference data sets or functions for processing user data. There isn’t a need to duplicate a reference data or processing functions for each user session, so declaring these types of items in the session-global scope is appropriate. However, I often find that beginning developers tend to declare variables in this scope because it is easy – you can access and use your variables and functions anywhere in the session.R file…
Some of the downsides to mis- or over- use of the global and session-global environments include:
- Environmental pollution (and usage collisions/conflicts)
- Data/Security exposure issues (all user sessions get these global values!)
- Memory cleanup issues
These three items can be extremely hard to track-down and fix in a large application, especially once you have multiple application users and/or multiple interconnected applications.
So consider using the most localized scope whenever possible: the session-local scope! This scope is limited to a single shiny application session. Unless your application does some very fancy footwork, it is possible for the user to create multiple sessions of an application, so this scope is limited by session, not just user. Within that application session anything changed, updated, used, etc. does not affect any other session – and this is usually what you want in your application! One user’s inputs, workflow, changes, etc. do not affect any other application user. This scope should be where the majority of non-reference objects and functions should be declared and used.
It is difficult to illustrate the adverse effects of misusing the global and session-global scopes without a number of simultaneous users of a shiny application. However these scopes apply no matter how simple, or complex, your shiny application. To illustrate where these scopes are I started with the familiar sample Observer shiny application (from RStudio) and made some modifications to illustrate scope in a simple manner. You can copy-paste the code into your own .R files and run it or download the three files using the button below (which are better spaced and well commented) to run the application. This application uses three variables scoped as in the discussion above and illustrated using the matching three colours below.
global.R
ui.R
session.R
You will notice that the code purposely changes the session-global variable from within the local session – so if you open up the application in multiple browser windows you can change the session-global variable and see how it does affect all sessions, not just the one open window. The session-local variable only affects the local session, and the overall-global variable is included to show you how declared items in “global.R” can be used in the application if desired.
Explore and change the sample app so that you can get a good feel for the different shiny scopes. If you can – open two different browsers or tabs with the application and see how your changes in one session affect the other sessions and vice-a-versa. A solid understanding of these three scopes in a shiny application will really help your development in the future be more professional, consistent, and solid!
Download the Example Scoping Shiny Code Here